Step 1: Single Cell Suspension Preparation
1. Sample Collection
2. Tissue mincing

3. Tissue preparation
4. Mechanical and enzymatic dissociation
5. Sample prepared
6. Single Cell Suspension
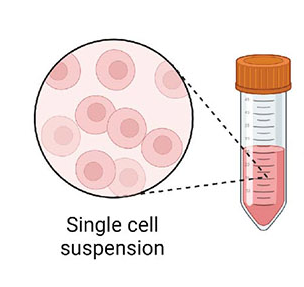
Spending time researching and learning how to create a quality single cell suspension BEFORE generating cDNA libraries with the 10x Chromium platform is essential.
A general rule of thumb is that if you are proficient in flow cytometry cell preparations you are prepared for the 10X Chromium step.
Preparation of single cell suspension from tissue samples or culture systems is the most important step in single-cell-RNA-seq.
Step 2: Microfluidics with 10x Chromium controller
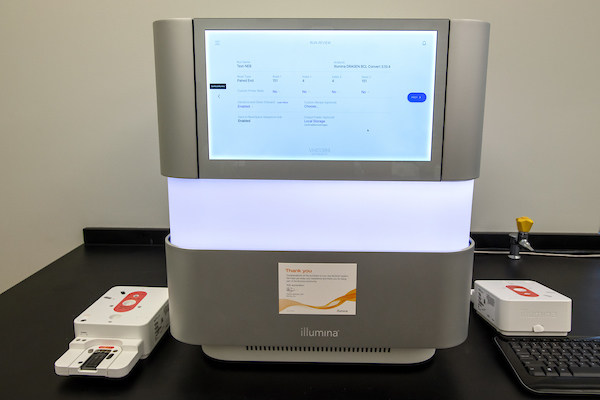
Step 3: cDNA library Preparation and Illumina Sequencing
Step 4: Sequence Alignment
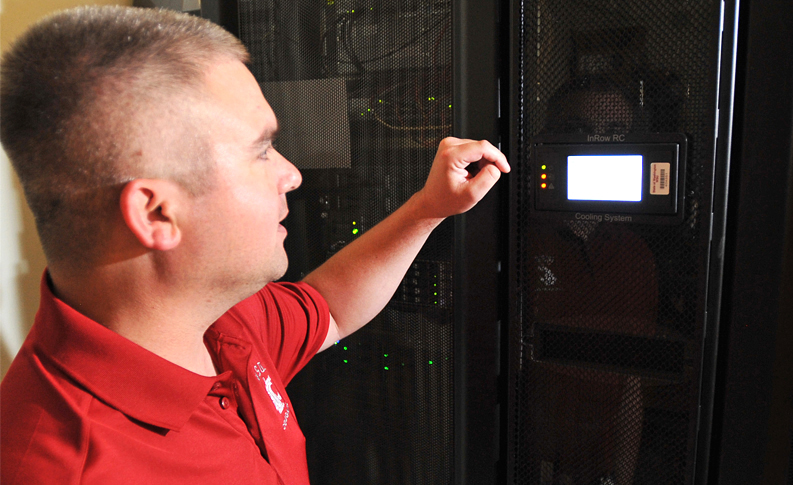

Sequence files called ‘FASTQs” will be generated by the Illumina sequencer that can be processed by using WSU’s high performance computer cluster called ‘Kamiak’, which has the 10x specific sequence aligner called ‘Cellranger’. Contact kamiak.support@wsu.edu to establish a user account and to receive instructions on how to align your FASTQs.
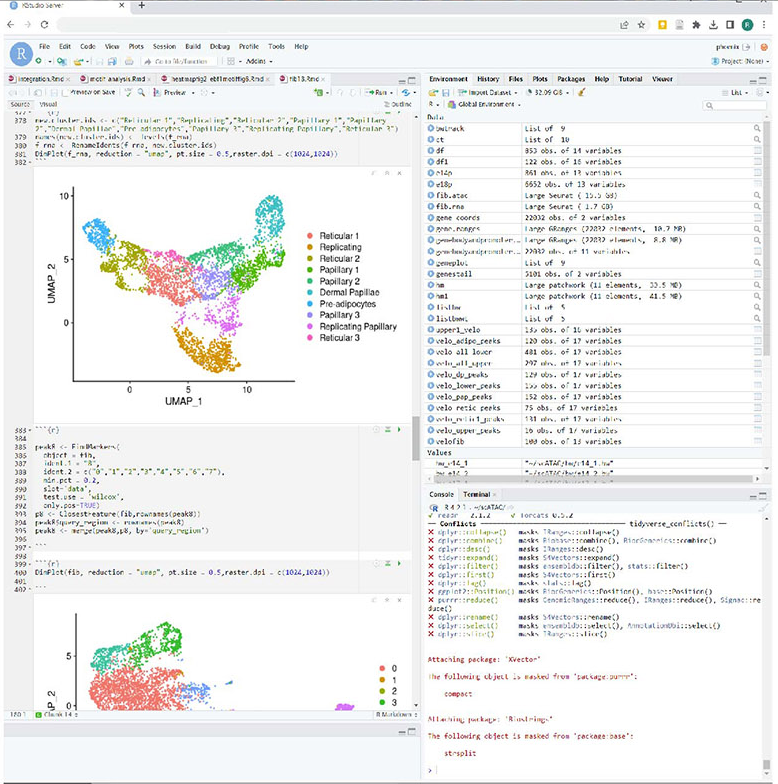
Step 5: Data Analysis
Once you have completed sequence alignment with CellRanger on Kamiak you can contact VIS to setup a cloud based server that will be ready for you to analyze the output files using Seurat on R.
Veterinary information systems
College of Veterinary Medicine Datahub: This is the CVM datahub that hosts webtools that are part of manuscript.
Technical Assistance or Service Inquiries:
509-335-0101 or help@vetmed.wsu.edu